
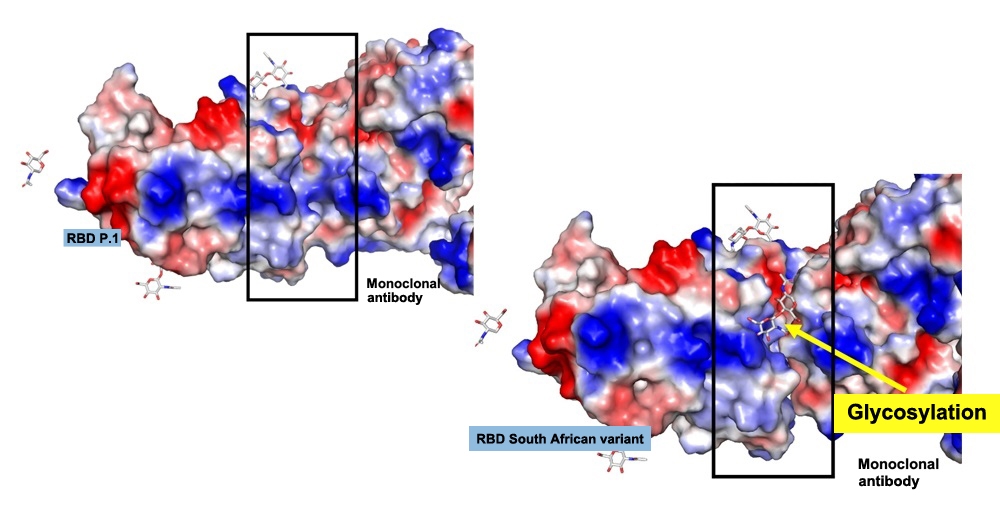
In the South African variant (B.1.351, right), the glycosylated site leaves a gap between the viral spike protein receptor and the antibody. In P.1 (left), the antibody binds more tightly to the virus (image: researchers’ archive)
A mutation in the variant’s spike protein is associated with glycosylation, which prevents antibodies from binding strongly to the virus. If the discovery is confirmed by further research, it could serve as a basis for future vaccines that are more effective against novel variants.
A mutation in the variant’s spike protein is associated with glycosylation, which prevents antibodies from binding strongly to the virus. If the discovery is confirmed by further research, it could serve as a basis for future vaccines that are more effective against novel variants.

In the South African variant (B.1.351, right), the glycosylated site leaves a gap between the viral spike protein receptor and the antibody. In P.1 (left), the antibody binds more tightly to the virus (image: researchers’ archive)
By André Julião | Agência FAPESP – A group of scientists led by researchers at the University of São Paulo’s Medical School (FM-USP) in Brazil believe they have found the mechanism that enables the South African variant of SARS-CoV-2, also known as B.1.351, to escape the antibodies produced during previous infections by the ancestral strain of the virus. If confirmed in further experiments, the discovery will pave the way to the development of vaccines that will be effective against both the variant first identified in South Africa and now present in Brazil, and the variant first identified in Manaus (P.1), as well as their predecessors.
An article reporting the results of the study is published on the preprint platform medRxiv and is being peer-reviewed.
The group performed computer simulations to study antibody recognition of the SARS-CoV-2 spike protein, which binds to the ACE-2 receptor on human cells in order to infect them. The findings suggest that a mutation in the docking tip of the South African variant’s spike (substituting the amino acid lysine for the amino acid asparagine) may entail its glycosylation, a biochemical phenomenon that changes the structure of the spike protein and prevents antibodies from binding to it. In P.1, lysine is replaced by threonine, which appears not to undergo glycosylation.
“In developing vaccines from now on, it will be necessary to choose the elements that will be most effective against the virus and its variants. Two of the three mutations in P.1 and B.1.351 are identical, so a vaccine that focuses on the South African variant could also be effective against P.1 and the ancestral virus, whereas a vaccine targeting P.1 and the ancestral strain will probably be less effective against the South African variant,” said Keity Souza Santos, a professor at FM-USP and corresponding author of the article.
The study was part of a project supported by FAPESP and led by Jorge Elias Kalil Filho, last author of the article. Kalil is a professor at FM-USP and heads the Immunology Laboratory at the Heart Institute (INCOR) of Hospital das Clínicas, the hospital complex run by FM-USP.
Kalil and his group are developing a COVID-19 vaccine with support from FAPESP and the Brazilian Innovation Agency (FINEP) (read more at: agencia.fapesp.br/32761).
Target
“Previous research by other groups failed to identify the specific region in which human antibodies couple with the RBD [receptor binding domain], the docking tip of the spike protein that slots into human cells. Only inferences have been possible until now. We used a technique that enabled us to locate a predominantly recognized region, which we call immunodominant. It’s precisely the site of one of the mutations in the Manaus and South African variants,” Santos said.
Having identified this region in the original virus, the research group – which comprises scientists affiliated with the University of São Paulo (USP), São Paulo State University (UNESP) and the Federal University of São Paulo (UNIFESP), as well as collaborators at the University of Salzburg in Austria – submitted the amino acid sequence to blood serum samples from 71 patients who had recovered from COVID-19 at Hospital das Clínicas early in the pandemic in Brazil. In 68% of the samples, the antibodies present in the serum were able to bind to P44, the most recognized peptide in the spike protein’s RBD.
The researchers performed computer simulations to understand how antibodies bind to this peptide. Data on the RBD in the two variants and the ancestral virus was correlated with data on the monoclonal antibody REGN10933, known to bind to the immunodominant region and currently in clinical trials for treatment of COVID-19. The software predicted the antibodies’ capacity to neutralize the virus.
Predicted neutralization was complete against the ancestral virus and fairly good against P.1, but poor against the South African variant (B.1.351), confirming the findings reported in Cell by US scientists shortly before the Brazilian researchers posted their own report.
For the group led by FM-USP, one of the variant’s mutations replaces lysine with asparagine, which is glycosylated, and this alteration probably accounts for the poorly predicted neutralization of the variant by antibodies deriving from the immune response to the ancestral virus. Glycosylation has been observed in the influenza virus, but not until now in SARS-CoV-2.
“In P.1 and B.1.351, the RBD mutation consists of only three different amino acids compared with the ancestral strain’s RBD,” Santos said. “Nevertheless, the change appears sufficient to make these two variants more transmissible. A vaccine that triggers the production of antibodies that attack the two mutations these variants have in common, plus the glycosylated amino acid in B.1.351, will probably be more effective.”
The group now plans to conduct in vitro experiments on samples of the peptide P44 to confirm whether antibodies really fail to bind to asparagine when it is glycosylated. In addition, they have obtained serum from recovered patients with P.1 and intend to find out whether their antibodies bind to P44.
The article “Immunodominant B cell epitope in SARS-CoV-2 RBD comprises a B.1.351 and P.1 mutation hotspot: implications for viral spread and antibody escape” is at: www.medrxiv.org/content/10.1101/2021.03.11.21253399v1.
Republish
The Agency FAPESP licenses news via Creative Commons (CC-BY-NC-ND) so that they can be republished free of charge and in a simple way by other digital or printed vehicles. Agência FAPESP must be credited as the source of the content being republished and the name of the reporter (if any) must be attributed. Using the HMTL button below allows compliance with these rules, detailed in Digital Republishing Policy FAPESP.




