

Understanding how microorganisms regulate the control and production of these enzymes is considered fundamental to the production of second-generation ethanol (photos: researcher's archive)
Understanding how microorganisms regulate the control and production of these enzymes is considered fundamental to the production of second-generation ethanol.
Understanding how microorganisms regulate the control and production of these enzymes is considered fundamental to the production of second-generation ethanol.
Understanding how microorganisms regulate the control and production of these enzymes is considered fundamental to the production of second-generation ethanol (photos: researcher's archive)
By Elton Alisson | Agência FAPESP – Production of second-generation (2G) ethanol from sugarcane requires enzymatic hydrolysis, in which enzymes from microorganisms act together to break down and convert the carbohydrates in sugarcane straw and bagasse into sugars capable of undergoing fermentation.
Understanding the genetic mechanisms that regulate the control and production of hydrolytic enzymes by microorganisms is considered fundamental to improving the technology used in this process.
Important knowledge of the different biological mechanisms behind the control and production of hydrolytic enzymes specifically by fungi has been garnered by a group of researchers at the University of Campinas (UNICAMP) in São Paulo State, Brazil, partnering with colleagues from the National Bioethanol Science & Technology Laboratory (CTBE), which belongs to the National Energy & Materials Research Center (CNPEM) in Campinas, and from Rio de Janeiro State University (UERJ).
Conducted as part of a project supported by FAPESP, the study was published in Scientific Reports.
“Our discoveries can contribute to the development of enzymes for inclusion in enzymatic cocktails used to produce 2G ethanol and other products,” said Anete Pereira de Souza, a professor at UNICAMP and principal investigator for the project, in an interview given to Agência FAPESP.
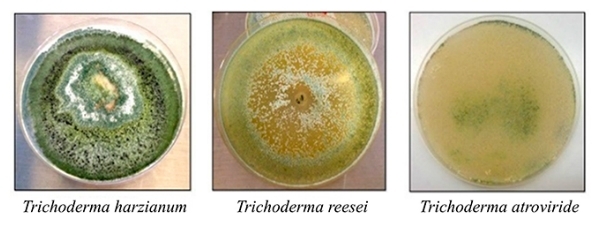
The researchers analyzed the genetic mechanisms involved in the secretion and expression of enzymes used by three species of fungus to degrade sugarcane. The species were Trichoderma harzianum, T. reesei and T. atroviride.
These fungi are frequently found in soil and growing on wood, bark and even other fungi, as well as many other substrates. They hydrolyze various kinds of carbohydrate, including the cellulose in sugarcane straw and bagasse, by means of enzymes present in their cell walls.
The researchers used several techniques in biotechnology and bioinformatics to find out whether the enzymes produced by the three Trichoderma species have similarities and differences that may enhance or limit their efficiency in breaking down biomass, as well as whether they behave synergistically during this process.
They first measured the activity levels of enzymes secreted by the three fungal species during fermentation of bagasse, pure cellulose and sugarcane glucose. To do this, they counted and analyzed the proteins present in these three different substrates at the height of the biodegradation process.
They then used a high-throughput RNA-sequencing technique called RNA-seq to identify the genes expressed. Through the use of bioinformatics tools, they compared the data and were able to pinpoint gene networks that are co-regulated by the three fungal species and could be essential for biomass breakdown by these microorganisms.
“We identified highly synergistic gene co-regulation networks involved in enzymatic degradation of sugarcane biomass by the three fungal species,” said Jaire Alves Ferreira Filho, who is studying for a PhD in genetics and molecular biology at UNICAMP and is one of the authors of the study.
High degree of synergy
The researchers identified 80 proteins and their respective genes shared by the three fungal species. They found 19 of these proteins in all three fungal species.
The 19 proteins and their respective genes are involved in the production and secretion of hydrolytic enzymes and are associated with different fungal mechanisms of biomass breakdown, the researchers explained.
The elucidation of genetic relationships between the sets of genes provides important information for the development of recombinant microorganisms with potential industrial applications, they added, while simultaneously contributing to the understanding of the synergistic reactions among enzymes.
“Our detailed description of these reactions will lead to significant advances. It provides a sound basis for the use of genetic information in the production of biofuels and countless biocompounds,” said Maria Augusta Crivelente Horta, first author of the article. Horta’s postdoctoral research was supported by a scholarship from FAPESP.
The Scientific Reports article “Network of proteins, enzymes and genes linked to biomass degradation shared by Trichoderma species” (doi: 10.1038/s41598-018-19671-w) by Maria Augusta Crivelente Horta, Jaire Alves Ferreira Filho, Natália Faraj Murad, Eidy de Oliveira Santos, Clelton Aparecido dos Santos, Juliano Sales Mendes, Marcelo Mendes Brandão, Sindelia Freitas Azzoni and Anete Pereira de Souza can be read at: nature.com/articles/s41598-018-19671-w.
Republish
The Agency FAPESP licenses news via Creative Commons (CC-BY-NC-ND) so that they can be republished free of charge and in a simple way by other digital or printed vehicles. Agência FAPESP must be credited as the source of the content being republished and the name of the reporter (if any) must be attributed. Using the HMTL button below allows compliance with these rules, detailed in Digital Republishing Policy FAPESP.





