

Cell culture trial shows interaction between cellular protein PCNA and SARS-CoV-2 protein M. Human protein PCNA (red) migrates from cell nucleus (blue) to cytoplasm in the presence of protein M (green) (credit: Orlando B. Scudero/ICB-USP)
Brazilian researchers have discovered that PCNA, a protein present in the nuclei of human cells, interacts with the SARS-CoV-2 membrane matrix protein M. In laboratory tests, inhibition of this mechanism using a drug reduced viral replication by 15%-20%.
Brazilian researchers have discovered that PCNA, a protein present in the nuclei of human cells, interacts with the SARS-CoV-2 membrane matrix protein M. In laboratory tests, inhibition of this mechanism using a drug reduced viral replication by 15%-20%.
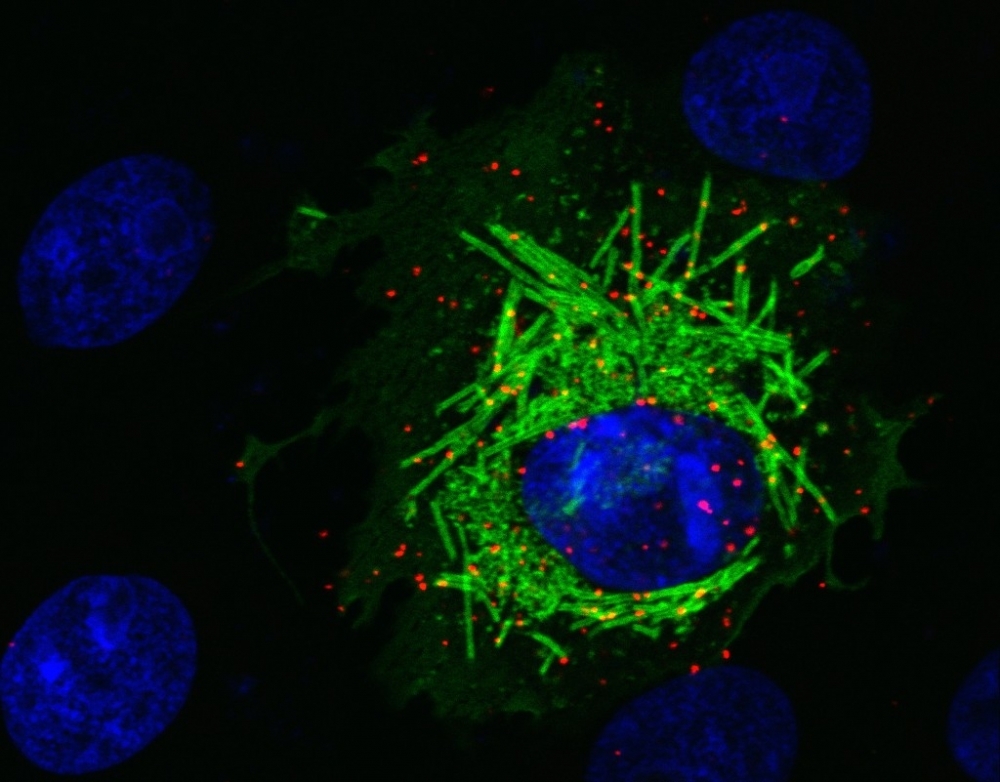
Cell culture trial shows interaction between cellular protein PCNA and SARS-CoV-2 protein M. Human protein PCNA (red) migrates from cell nucleus (blue) to cytoplasm in the presence of protein M (green) (credit: Orlando B. Scudero/ICB-USP)
By André Julião | Agência FAPESP – An article published in the journal Frontiers in Cellular and Infection Microbiology reports a study by researchers at the State University of Campinas (UNICAMP) and the University of São Paulo (USP) in Brazil showing how a human protein interacts with a SARS-CoV-2 protein, and describing one of the ways the virus that causes COVID-19 recruits cells to replicate.
In laboratory tests, the researchers inhibited interaction between the molecules using a drug and thereby reduced viral replication by 15%-20%. They expect their findings to contribute to the development of treatments for COVID-19.
“The human protein known as PCNA [proliferating cell nuclear antigen] interacts with the SARS-CoV-2 protein M [matrix], one of the molecules that make up the virus’s membrane and give it shape. The discovery itself shows one of the ways the pathogen manipulates cell function for its life cycle to proceed,” said Fernando Moreira Simabuco, a professor at UNICAMP’s School of Applied Sciences (FCA) in Limeira and principal investigator for the study, which was supported by FAPESP.
The group used a range of in vitro techniques to investigate how the presence of the viral protein M in the organism makes PCNA, a protein involved in DNA repair, migrate from the cell nucleus, where it is normally found, to the cytoplasm, a cellular region containing organelles responsible for important cell functions.
According to the researchers, this migration shows that the viral and human proteins interact, a conclusion corroborated by other methods, such as use of compounds to inhibit migration of proteins from the nucleus to the cytoplasm. In cells treated with both a specific compound for PCNA and another that inhibits migration of different proteins including PCNA, viral replication was reduced by between 15% and 20% compared with untreated cells.
“If we’d been thinking about treatment, perhaps this reduction wouldn’t have been significant, but our main aim was to demonstrate the interaction and show that it could be a future therapeutic target,” Simabuco said.
In collaboration with researchers in the Pathology Department of USP’s Medical School, they analyzed samples of lung tissue obtained during autopsies of deceased COVID-19 patients (more at: agencia.fapesp.br/32955/).
Expression of PCNA was found to be above normal in these samples, as was expression of the protein gamaH2AX, a marker of DNA damage, reinforcing the results.
“This finding may point to yet another consequence of infection by the virus,” Simabuco said.
The first author of the article is Érika Pereira Zambalde, a postdoctoral researcher at FCA-UNICAMP under Simabuco’s supervision.
Protein news
The protein M is anchored, with proteins E and S, in the membrane that envelops SARS-CoV-2, and is the most abundant of its four main structural proteins, called structural because they give it shape. For this reason, it has been considered a potential target for medications and vaccines.
S, the viral spike protein, is well-known because it binds to the ACE receptor in human cells, a role that has made it the target for most current COVID-19 vaccines.
The human protein PCNA is widely studied in the context of cancer research, as exemplified by a project conducted by Simabuco at FCA-UNICAMP. Little is known about the role of PCNA in viral infections, however.
The recently published article, therefore, offers a way forward for further research on this interaction between SARS-CoV-2 and PCNA, facilitating the development of therapies. A next step would be validation of the discoveries in animal models, although this has not yet been programmed.
Some of the experiments were conducted at the Laboratory of Emerging Virus Studies (LEVE) headed by José Luiz Proença Módena at UNICAMP’s Biology Institute (IB), with FAPESP’s support.
Research groups led by Armando Morais Ventura, a professor at USP’s Biomedical Sciences Institute (ICB), and Henrique Marques-Souza, a professor at IB-UNICAMP, collaborated in the study.
The article “Characterization of the interaction between SARS-CoV-2 membrane protein (M) and proliferating cell nuclear antigen (PCNA) as a potential therapeutic target” is at: www.frontiersin.org/articles/10.3389/fcimb.2022.849017.
Republish
The Agency FAPESP licenses news via Creative Commons (CC-BY-NC-ND) so that they can be republished free of charge and in a simple way by other digital or printed vehicles. Agência FAPESP must be credited as the source of the content being republished and the name of the reporter (if any) must be attributed. Using the HMTL button below allows compliance with these rules, detailed in Digital Republishing Policy FAPESP.





