
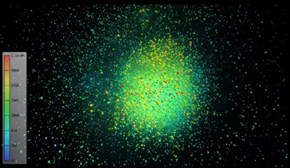
A gene-gene interaction network in the CA3 region of the human hippocampus: the color scale on the left indicates the number of connections between one gene and another
Researchers use a new strategy to study the brains of people with a type of drug-resistant epilepsy and are also using the method to understand the development of the thymus.
Researchers use a new strategy to study the brains of people with a type of drug-resistant epilepsy and are also using the method to understand the development of the thymus.

A gene-gene interaction network in the CA3 region of the human hippocampus: the color scale on the left indicates the number of connections between one gene and another
By Karina Toledo
Agência FAPESP – Discovering how the genes in particular tissues in the human body communicate and the changes in the gene interaction network that occur when a person is ill allows scientists to better understand the molecular mechanism of these infirmities and how to identify therapeutic targets for the development of new drugs.
Researchers from Universidade de São Paulo (USP) are using this strategy to study the brains of people with a type of drug-resistant epilepsy. These researchers are also using the method to understand the development of the thymus, an organ of major importance for the immune system, with the future objective of discovering how autoimmune and immunodeficiency diseases develop.
“We are using a tool in field of Genomics that emerged in Physics some time ago: the analysis of complex networks. This allows for the mapping of the most important genes and those that have more connections to other genes,” explained Carlos Alberto Moreira-Filho of the USP Medical School (FMUSP).
The analysis is conducted with millimeter-sized samples of the tissue being studied. Scientists extract the messenger RNA found in the fragment and measure which genes are the most or least expressed at the location using statistical analysis.
“Our genes are the same in any part of the body. What distinguishes a retina cell from an epithelial cell or gastric mucosal cell is the set of genes that is expressed and the interaction network among them. Through pair by pair statistical analysis, it is possible to perceive when the expression of a gene increases or decreases and the other components that rise or fall with it. That is how we map an interaction network,” stated Moreira-Filho.
This analysis allows for the identification of two types of key genes in tissues: the HUBs – those that have a large number of connections with other genes – and the VIPs – those that do not have many connections but function as a bridge between the HUB-type genes.
“Identifying the genes that are VIP genes and those that are HUB genes is not mere statistical curiosity. It is extremely important in terms of biological function. HUB genes are related to an important metabolic pathway, and VIP genes are responsible for uniting two or more metabolic pathways in a process,” explained Moreira-Filho.
According to Moreira-Filho, the software developed for these analyses could be used to study disease in any part of the body. Such software also could serve as a tool for functional genomic studies on microorganisms, plants and animals, selecting useful information for research that is directed toward, for example, genetic improvement.
The study has been conducted under the auspices of the project “Models and methods of e-Science for life and agricultural sciences,” coordinated by Roberto Marcondes Cesar Junior of USP’s Mathematics and Statistics Institute (IME-USP) and including collaboration from the group led by Luciano Fontoura da Costa at São Carlos Institute of Physics (IFSC-USP).
The project is funded by FAPESP and the National Council of Scientific and Technological Development (CNPq) through the Program for the Support of Excellence Centers (Pronex).
Synapse
The analyses on epilepsy, however, began even before the Pronex funding under the auspices of a Thematic Project coordinated by Moreira-Filho. The objective was to understand why some patients with a more common form of the disease – known as mesial temporal lobe epilepsy – do not respond to drug treatment.
“We estimate that a third of the individuals affected by this type of epilepsy in the world are medically refractory to the existing drugs, totaling approximately 10 million people,” Moreira-Filho explained.
These patients can have several crises per week, which can compromise their quality of life in addition to increasing their risk of sudden death. Currently, the only option in these cases is to surgically remove the affected part of hippocampus – a difficult, expensive and invasive procedure to which few have access.
“The ideal [scenario] is finding a drug solution. However, to do this, we need to better understand the mechanism of the disease, and one of the most interesting ways to accomplish this is to study the genetic interaction network,” affirmed Moreira-Filho.
At first, researchers used simpler software that was capable of solely mapping the network of differentially expressed genes – those that are expressed differently due to a stimulus from the environment, such as fever or trauma. At the time, the network had between 200 and 400 genes. Currently, thanks to more complex software, there are roughly 15,000 genes, which corresponds to practically all the genes expressed in the hippocampal region.
“Today, we know for sure that being differentially expressed is not the only reason why a gene is relevant in a disease. Sometimes, this is a consequence of a gene relationship pattern that has changed. We realize that the differentially expressed genes are an element of disturbance in the network. However, to know where to intervene, we must understand the entire network,” explained Moreira-Filho.
The initial investigations into the mesial temporal lobe epilepsy area resulted in a study published in the journal PLoS One. In the article, the researchers showed that the region acquires a different molecular profile, depending on the stimulus that sets off the disease.
“There is a form of the disease that results from febrile seizures and another from other types of stimuli. These produce different profiles from a molecular point of view, and this is important for identifying therapeutic targets. We have already identified some candidate genes for drug studies in vitro and with animal models,” Moreira-Filho stated.
The research with fragments of the thymus is still in the initial stage, but the scientists were able to detect that the gene expression patterns in this organ underwent major alterations at 6 months of age and again at 12 months. The preliminary results were presented at the 2nd São Paulo School of Advanced Sciences in Primary Immunodeficiency Diseases (ESPCA-PID), held from March 3-8, 2013.
“The thymus is responsible for the development of central tolerance, or rather, it teaches the defense cells not to attack the antigens of the organism. The organ is large in newborns and shrinks over time, but the tolerance remains. We want to understand what happens that is so spectacular during the first year of life,” affirmed Moreira-Filho.
Republish
The Agency FAPESP licenses news via Creative Commons (CC-BY-NC-ND) so that they can be republished free of charge and in a simple way by other digital or printed vehicles. Agência FAPESP must be credited as the source of the content being republished and the name of the reporter (if any) must be attributed. Using the HMTL button below allows compliance with these rules, detailed in Digital Republishing Policy FAPESP.





