
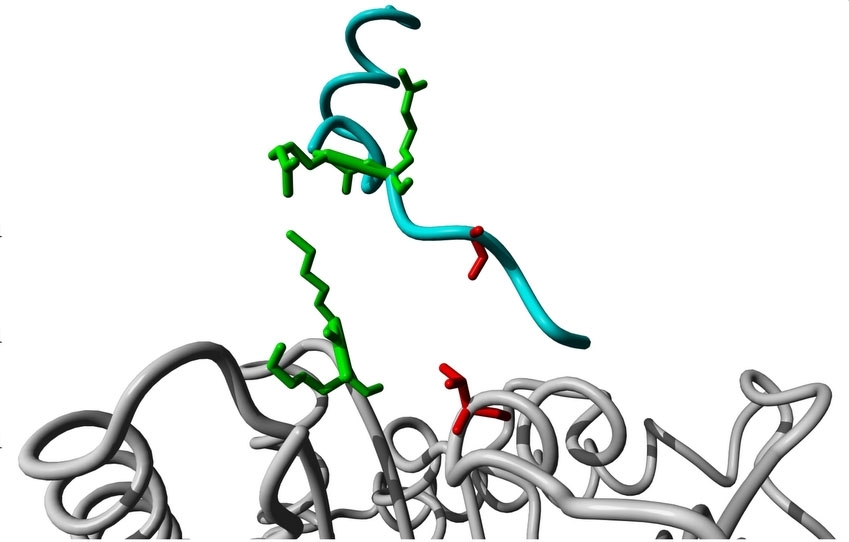
A research study featured on the cover of the journal Science Signaling explains the interplay of a chemical reaction known as phosphorylation, which is catalyzed by kinases (image: Deborah Schechtman)
A research study featured on the cover of the journal Science Signaling explains the interplay of a chemical reaction known as phosphorylation, which is catalyzed by kinases.
A research study featured on the cover of the journal Science Signaling explains the interplay of a chemical reaction known as phosphorylation, which is catalyzed by kinases.

A research study featured on the cover of the journal Science Signaling explains the interplay of a chemical reaction known as phosphorylation, which is catalyzed by kinases (image: Deborah Schechtman)
By Karina Toledo
Agência FAPESP – Kinases are enzymes that act as a type of protein ‘on/off switch’ and, in this way, regulate important processes in the body such as cell division, proliferation and differentiation.
Any problems in this signaling event may result in pathologies such as cancer, inflammation or cardiovascular disease. For this reason, nearly 25% of efforts by the pharmaceutical industry target the development of compounds that are capable of modulating the activity of these enzymes.
A study carried out by Brazilian researchers that was featured on the cover of the journal Science Signaling may make it easier to identify the target protein (or substrate) of a particular kinase, which is no easy task considering that there are over 500 different enzymes of this type in the human genome.
“When we’re able to identify the substrate of a kinase, we’re able not only to better understand its function in the body but also to design compounds capable of more selectively modeling its action, causing fewer side effects,” said Deborah Schechtman, professor in the Chemistry Institute of the University of São Paulo (USP) and coordinator of the FAPESP-funded research study.
Schechtman explained that kinases are responsible for a chemical reaction known as phosphorylation, which involves adding a phosphate group of high-energy molecules such as ATP (adenosine triphosphate) to the target protein. This reaction may, depending on the case, activate or deactivate the protein or signal for the molecule to be degraded.
However, for the interaction to take place, the enzyme and the substrate need to fit together similarly to the pieces of a jigsaw puzzle. The target protein has to have a specific sequence of amino acids, technically called a “consensus sequence,” that is capable of being recognized by the kinase.
“Previous studies have looked for this consensus sequence in the linear structure of the protein [on the chain of amino acids chemically linked to one another]. Our work showed that in some cases, the sequences can only be found when looking at the three-dimensional structure of the molecule. This is a shift in paradigm,” said Schechtman.
For the protein to perform its function inside the cell, the chain of amino acids that makes it up must be folded into a specific three-dimensional shape through a process known as protein folding.
When this occurs, amino acids that were once far from each other in the primary structure become closer and, in some cases, form what Schechtman’s group referred to as a consensus motif, which may also be recognized by a kinase.
“We determined that the so-called anchor residues, which form the consensus motif that will be recognized by the kinase, are important for the interaction between enzymes and substrates and may be targets for developing inhibitory drugs with highly specific actions,” Schechtman said.
According to the researcher, the inhibitors available in the pharmaceutical arsenal today impede the transfer of the phosphate group to the proteins but indiscriminately affect the kinases of the body, which may hinder physiological processes.
Tubulin
The line of investigation that led to the discovery published in the journal Science Signaling began in an earlier FAPESP-funded project in which Schechtman attempted to discover substrates of the kinase PKC (protein kinase C), which is involved in the proliferation and differentiation of embryonic stem cells.
“I discovered that PKC interacted with a protein called tubulin, which takes part in the process of cell division. We then started to investigate that substrate. Using a technique known as mass spectrometry, we identified which amino acid residue of tubulin that PKC interacted with and realized that there was no consensus sequence around it that the kinase recognized,” the researcher explained.
Working together with the research group led by Paulo Sergio Lopes de Oliveira from the Brazilian Biosciences National Laboratory (LNBio), Schechtman decided to study the three-dimensional structure of tubulin and found what she called the consensus motif.
“To confirm it, we produced some mutated versions of tubulin with different anchor residues and observed that recognition by the kinase was greatly compromised,” Schechtman explained.
The next step was to prove that the consensus motif could also be found in other proteins. For this, the researchers developed a model.
“Because we needed to prove a concept and there are few known substrates for PKC, we conducted the modeling using PKA (protein kinase A), another well-studied kinase. That way, the sampling would be larger,” Oliveira said.
The researchers then analyzed a database of experimentally detected PKA phosphorylation sites, compared them with information from a database of three-dimensional protein structures determined using crystallography, and used modeling to assess how the motifs would line up in three dimensions.
“We modeled more than a thousand PKA substrates and saw that in 27 cases, the anchor residues presented only the consensus motifs,” Oliveira said.
The results of the modeling also suggest that in order for the protein to be recognized by the kinase, both the order of the amino acids in the structure and the distance between them are determining factors, similarly to parts of a jigsaw puzzle.
Schechtman’s group continues to study tubulin in an effort to understand the effect of phosphorylation during cell division. The researcher’s goal is to better understand the molecular processes involved in embryonic stem cell differentiation, which is essential for their safe use in cell therapy.
Republish
The Agency FAPESP licenses news via Creative Commons (CC-BY-NC-ND) so that they can be republished free of charge and in a simple way by other digital or printed vehicles. Agência FAPESP must be credited as the source of the content being republished and the name of the reporter (if any) must be attributed. Using the HMTL button below allows compliance with these rules, detailed in Digital Republishing Policy FAPESP.




