
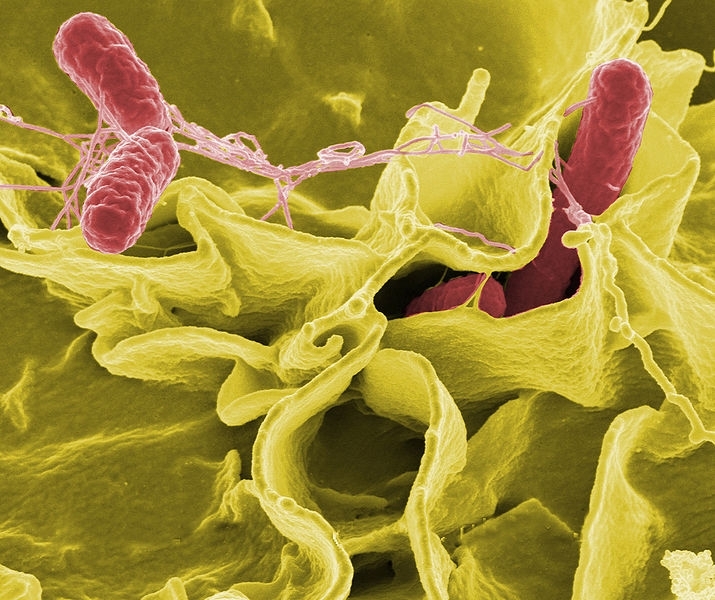
Project presented at FAPESP Week London identifies genes that allow bacteria to survive in the digestive tract of poultry and thus infect humans (image: Wikimedia Commons)
Project presented at FAPESP Week London identifies genes that allow bacteria to survive in the digestive tract of poultry and thus infect humans.
Project presented at FAPESP Week London identifies genes that allow bacteria to survive in the digestive tract of poultry and thus infect humans.

Project presented at FAPESP Week London identifies genes that allow bacteria to survive in the digestive tract of poultry and thus infect humans (image: Wikimedia Commons)
By André Julião | Agência FAPESP – A group of researchers from the School of Agricultural and Veterinary Sciences, at the State University of São Paulo (FCAV-UNESP) in Jaboticabal, are investigating genes that are important for the survival of the bacterial species Salmonella in the intestinal tract of poultry. The goal is to prevent foodborne infections in humans.
There are more than 2,600 known serotypes of Salmonella. Some are responsible for many cases of infection in animals and humans. The presence of certain serotypes in Brazilian poultry products has caused the European Community to block containers exported by Brazil. European law is quite restrictive when it comes to the presence of these bacteria.
“Salmonella easily colonizes the digestive tract of poultry and may or may not cause disease. Even when they do not affect the chickens themselves, they are able to infect humans who eat the chicken,” said Angelo Berchieri Junior, a professor at FCAV-UNESP during a session at FAPESP Week London held February 11-12, 2019.
Berchieri is responsible for a FAPESP-funded Thematic Project that will test the effects of deleting ttrA and pduA genes from three serotypes of Salmonella: Salmonella Enteritidis, S. Typhimurium and S. Heidelberg.
“We selected these three serotypes because they are frequently found in poultry and may cause foodborne infections in humans,” said Berchieri.
The researcher explained that of the three, S. Heidelberg is the least common among humans. However, it was found in the Brazilian poultry shipments that were not accepted in Europe. “The serotype is widespread in Brazil and could compromise Brazilian exports,” he said.
Brazilian law specifically restricts serotypes Enteritidis and Typhimurium. However, depending on the country of import, other serotypes of Salmonella in exported poultry products may also come under restrictions.
Also taking part in the project is post-doc Mauro de Mesquita Souza Saraiva and biologist Gabriele Tostes Gricio, recipient of a technical training scholarship.
To determine which genes cause the bacteria to become resistant to the immune system of poultry, the researchers infected one group of chicks with a wild type bacterium [without genetic modification] and the other group with Salmonella whose ttrA or pduA genes had been deleted.
Then they compared the two groups for the presence of bacteria in feces, the cecum [a pouch that forms the first part of the large intestine], the liver and the spleen.
By identifying the genes that allow the bacteria to survive, the researcher is able to generate mutant forms that may be used as a vaccine. When the immune system comes in contact with a variety that does not kill the animal, but survives in the organism for a period of time, that animal establishes an immunological memory. In the event the animal is later exposed to a harmful version of the bacteria its defenses will be ready to attack it.
“Understanding and combating the serotypes that may infect poultry is therefore critical for the health of consumers and Brazil’s trade balance,” said Berchieri.
Read more about FAPESP Week London at: www.fapesp.br/week2019/london/.
Republish
The Agency FAPESP licenses news via Creative Commons (CC-BY-NC-ND) so that they can be republished free of charge and in a simple way by other digital or printed vehicles. Agência FAPESP must be credited as the source of the content being republished and the name of the reporter (if any) must be attributed. Using the HMTL button below allows compliance with these rules, detailed in Digital Republishing Policy FAPESP.




