Diário de Campo – Capítulo 7: Expedição termina com mais de 130 espécies de peixes coletadas
Último episódio da série de reportagens faz um balanço sobre a viagem de um grupo de pesquisadores do Museu de Zoologia da USP que, ao longo de duas semanas, percorreu os rios Negro, Preto e Jauaperi, nos Estados do Amazonas e de Roraima
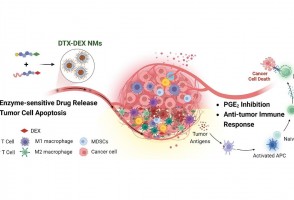
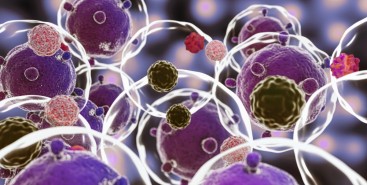
Cientistas do Brasil e da Índia criam tratamento promissor contra tumores sólidos
Em testes com animais, nanopartículas contendo substâncias já aprovadas para uso humano reduziram a inflamação no microambiente biológico em que cânceres desse tipo se instalam e vicejam, facilitando a ação do sistema imune
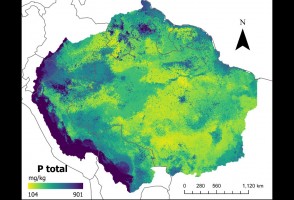
Mapas desenvolvidos com inteligência artificial confirmam baixos níveis de fósforo no solo da Amazônia
Pesquisa traz nova metodologia para descrever quantidade do mineral, que tem importância no ciclo de crescimento da vegetação e pode afetar a resposta da floresta às mudanças climáticas

Terceira Conferência FAPESP de 2024 aborda a formação de médicos-cientistas
Convidado é o professor da Universidade de Chicago Antonio Bianco, cuja pesquisa enfoca a ação dos hormônios tireoidianos
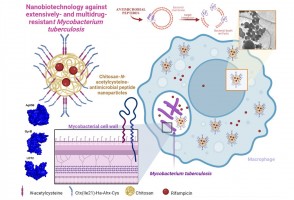
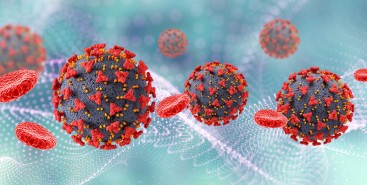
Pesquisadores da Unesp testaram a ação de partículas carregadas de antibióticos e outros compostos antimicrobianos em células infectadas pela bactéria causadora da doença. Resultados indicam que a estratégia é capaz de reverter o quadro de resistência aos medicamentos
Estudo focado na atenção primária busca melhorar diagnóstico e manejo da doença pulmonar obstrutiva crônica
Projeto conduzido por pesquisadores da USP e da Universidade de Birmingham foi um dos 15 apresentados em evento que celebrou a parceria entre a instituição britânica e a FAPESP. Ocasião também marcou o lançamento do University of Birmingham Brazil Institute, cuja missão é estreitar os laços de pesquisa com o Brasil
Chamada com UE apoiará pesquisas em ciência e engenharia de materiais
FAPESP lança edital em parceria com a M-ERA.NET, rede financiada pela União Europeia; pré-propostas serão recebidas até 14 de maio
OPORTUNIDADES FAPESP
Doutorado em biologia estrutural
CEPID B3/IQ-USP
Inscrições até 30/04/2024
Doutorado e mestrado em física da matéria condensada
IFSC-USP
Inscrições até 30/04/2024
PD em visão computacional
SemeAr/Embrapa Agricultura Digital
Inscrições até 30/04/2024
TT-4 em ecologia e conservação
FFCLRP-USP
Inscrições até 30/04/2024
PD em materiais magnéticos
IF-USP
Inscrições até 30/04/2024
Terceira Conferência FAPESP de 2024 aborda a formação de médicos-cientistas
Chamada com UE apoiará pesquisas em ciência e engenharia de materiais
Livro destaca a relevância das mulheres na construção do pensamento geográfico
Registro de produtos para controle biológico de pragas ultrapassa o de agroquímicos no Brasil
Em palestra a empresários, ganhadores do Prêmio Nobel destacam a importância da pesquisa básica
FAPESP recebe propostas para novas Escolas de Ciência Avançada até 3 de maio
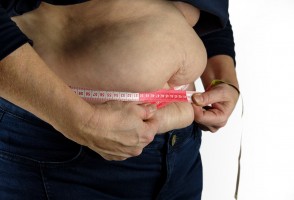
Obesidade abdominal associada à fraqueza muscular é condição que mais eleva risco de síndrome metabólica
Capítulo 6: Viagem ao rio Negro marcou reencontro após expedições pioneiras nos anos 1990

Uma das artes de pesca utilizadas para coletar peixes-elétricos na Expedição DEGy Rio Negro foi empregada pela primeira vez em larga escala em água doce no projeto Calhamazon, que reuniu pesquisadores do Brasil e dos Estados Unidos entre 1993 e 1996
Capítulo 5: Acidente com raia não atrapalha o ritmo das coletas no Jauaperi

Baixa gravidade da lesão, atendimento médico rápido e cuidados adequados fizeram com que pesquisador pudesse voltar aos trabalhos no mesmo dia em que foi ferroado por peixe peçonhento. Na Amazônia, casos muitas vezes se agravam por carência de assistência especializada
Capítulo 4: Sarapós se destacam pela diversidade de ambientes que ocupam

Os mais abundantes peixes-elétricos estão presentes desde o fundo dos grandes rios até os igarapés, onde podem se enterrar na areia ou se confundir com o folhiço. Em duas semanas, expedição na bacia do rio Negro coletou 27 espécies do grupo
Vídeos
Diário de Campo - Rio Negro
O Legado Suíço-Brasileiro na Amazônia: Arte, Ciência e Sustentabilidade
02/03/2024 a 30/04/2024
Introdução à programação com Python para alunas do ensino médio ou concluintes
30/03/2024 a 27/04/2024
Minicursos para estudantes do ensino médio 2024
06/04/2024 a 08/06/2024
Treinamento de plataformas e bases de dados científicas
09/04/2024 a 30/04/2024
14ª Semana da Computação
22/04/2024 a 26/04/2024
Chamada de Propostas para Centros de Pesquisa em Inteligência Artificial Aplicada à Saúde
24/04/2024 a 24/04/2024
Pulverização de lavouras vai ganhar aeronave especializada
Startup apoiada pelo PIPE-FAPESP está desenvolvendo helicóptero autônomo para pulverizar plantações em terreno íngreme
Teste irá permitir detecção pouco invasiva de micrometástases
Tecnologia pode possibilitar produção de enzimas no Brasil
Pulverização de lavouras vai ganhar aeronave especializada
Startup apoiada pelo PIPE-FAPESP está desenvolvendo helicóptero autônomo para pulverizar plantações em terreno íngreme
Chamadas FAPESP
Comunicar Ciência
Prazo: 22/01
Belmont Forum Climate, Environment, and Health
Prazo: Jan 2024
PIPE Start FAPESP-Sebrae: iniciando a jornada empreendedora de base tecnológica
Prazo: 18/03
Centros de Pesquisa em Inteligência Artificial Aplicada à Saúde
Prazo: 18/03
Fundação Nacional de Ciência da Suíça
Prazo: 22/03
Apoio a pesquisa em citricultura
Prazo: 31/03
Expedições Científicas Amazônia+10
Prazo: 29/04